AutozygosityMapper is primarily aimed at VCF files, we recommend using HomozygosityMapper to analyse SNP arrays.
FAQs: Click here to read the frequently asked questions
AutozygosityMapper can be used free of charge (even for commercial purposes) and without a login.
This software is being further developed. You might occasionally discover new features that haven't yet made it into the manual. We apologise in advance if this causes confusion, but we want to make new features available as soon as possible.
While we could import a variety of SNP chips and provide the infrastructure to upload genotypes created with them, we could not test all of them - simple due to the lack of the genotypes. If you cannot upload your data or want other species or chips to be included, please send an email to dominik.seelow (at) charite.de.
You can use AutozygosityMapper without a personal account. However, with a user profile, access to your data will be facilitated and you can also change the access restrictions (only you, everyone, selected co-operation partners) at any time.
It is not possible to choose a user name that is already in use, AutozygosityMapper will raise an error in
such cases.
You should always enter a valid e-mail address because we will inform you in advance when major changes to the
website are
planned or when we have to archive old projects due to disk space limitations.
If you forgot your password, you have the possibility to click on I
forgot my credentials! next to the help button on the login page. Afterwards you can enter the e-mail
address which you registered your account with, so that your credentials can be sent to you.
After you have created a password, you will be automatically logged in with these data. If you return to
AutozygosityMapper, please click on the login link on the homepage and enter your credentials.
If you want to upload, analyse, or view genotypes of any non-human species or want to switch to another species, simply click on the correct species in the panel on the right top of the homepage.
So far, AutozygosityMapper covers the following species:
 |
 |

|
|
| Humans Homo sapiens |
Cattle Bos taurus |
Dogs Canis lupus |
Horses Equus ferus caballus |

|
 |

|
|
| Mice Mus musculus |
Rats Rattus norvegicus |
Sheep Ovis aries |
Please see contributions for copy rights.
Genotypes from VCF files and the following chips can be imported:
Humans
|
Cattle
|
Mice
|
If your specied or chip is not listed here, please send us an email.
To import the genotypes, specify a new project name under new project name.
You can also decide whether or not you want to make the project public (checkbox access restriction).
If you are logged in with a personal account, this setting can be changed at any time and you can also share
your data with selected
collaborators (via the 'grant access to your data' link on the homepage). If you upload your data as guest, you
will receive special
URLs for access-restricted data; the data will be invisible to anyone else. Without the URL, you will not be
able to access your data again.
Also, it is not possible to use URL-protected genotypes in GeneDistiller to rank candidate genes by their
homozyosity scores.
Please note that data uploaded as guest will be deleted after two month to save disk space.
We hence recommend to create an account, especially if
you want to
keep your data private. Personal accounts are completely free.
After choosing a project name, you have to select wether you want to upload a 'VCF' or a genotyping chip (genotypes from VCF file or SNP chip). After choosing one of the options, more fields will appear. In case you choose the VCF file option, you have the possibility to set a minimum coverage and point to the genotype file (genotype file) to eventually submit the data. In case you choose the SNP chip option, you can select the respective SNP chip from a drop down menu (choose SNP chip) and afterwards point to the genotype file (genotype file) to eventually submit the data.
SNP genotype files must be tabular with the samples as columns and the SNPs as rows, they can (or rather should) also be zipped or gzipped. Lines starting with the number sign (#) will be ignored. Sample files to demonstrate the allowed formats can be found here. The genotypes must be written in standard Affymetrix or Illumina notation, i.e. (NoCall, AA, AB, BB or -1, 0, 1, 2 for Affymetrix and --, AA, AB, BB for Illumina, respectively).
AutozygosityMapper is primarily aimed at VCF files, we recommend using HomozygosityMapper instead.
VCF genotype files must combine the genotypes of all samples in one file (see below). A short description and a sample VCF file
can be obtained from the file formats page.
All variations from the RefSeq in your file will be used.
You can also set a coverage threshold (minimum coverage). Genotypes with a
coverage
below this value will be treated as no calls. Please note that the DP flag must
be included
in the FORMAT string (not only in INFO!), unless you
set the minimum coverage
value in the upload interface to 0. Without the DP flag in FORMAT it is impossible to
exclude
genotypes with a low coverage because the DP information in INFO aggegrates the
coverage over all samples!
If you want to combine different VCF files, you should create a joint file from the original BAM files. Using
bcftools,
this can be achieved by
bcftools mpileup -Ou -a DP -f reference.fa FILE1.bam FILE2.bam | bcftools call -mv -Ov -o VCFFILE.vcf # reference genome: reference.fa # output VCF file: VCFFILE.vcfPlease read the bcftools to find the appropriate settings for your data.
The data import may take seconds or even half an hour, depending on the size of the file, the upload speed and the working load of our server.
When the upload has been finished, you can proceed to the next step.
Mixing of arrays with different markers per sample is possible. However, AutozygosityMapper only
considers markers that
were genotyped for a sample. That means, that if you mix different arrays you will automatically obtain lower
homozygosity scores
for samples with fewer genotypes.
We believe that any other solution (using block length in megabases instead of SNP numbers, treating non-typed
markers as unknown
genotypes, or use only markers available in all samples) would merely complicate the application and make it
less performant in more
realistic scenarios. If, however, you are in such a situation, please drop us an e-mail. At attempt to solve
this might be worthwile,
challenging and exciting... :-)
After the genotypes were imported, you can start the homozygosity mapping. First, you must select the project you want to analyse (drop-down menu project) and provide a new name for the analysis (analysis name). You can specify further details under analysis description which will be displayed together with the analyses' names to indicate specific models etc. which might be too complex to include them in the name.
Under cases and controls, you can specify the samples that should be regarded as cases or controls, respectively. Cases are, of course, obligatory. Please note that sample IDs must be written exactly as in the genotype file; AutozygosityMapper will, however, complain, if it cannot find a sample.
If your cases are from a single family or if you are absolutely sure that there is no genetic heterogeneity among the samples you plan to analyse as cases, you can tick the require autozgysotiy box: The case samples must share the same homozygous genotype. In the text field exclude homozygous stretches in controls >= below, you can define a threshold for the length of homozygous blocks in controls. Whenever these exceed this value, the region will be excluded. The suggested values (printed next to the input field) are relatively high to reduce the risk of false negatives.
Another parameter to be set is the maximum block length (limit block length). If this field is left empty, the standard setting for chips will be used (6 mbp). If you don't want to limit the block length, specify a negative value. While the default settings are suitable for most analyses, after a visual inspection of the results you may decide to re-analyse your data according to the degree of homozygous blocks in your data. A small value might be appropriate if you want to find a founder mutation in a very large sample consisting of individuals from non-related families.
The next parameter to define would be the minimum number of variants. If you leave the field empty, the default value 10 will be used in case of vcf files and 4 in case of snp files.
Only count blocks longer than x . This parameter is an experimental way to fine-tune the block length settings. This is a lower limit of the block length; i.e. only homozygous stretches longer than n bases are be included in the score. This switch is helpful when you expect a large degree of genetic heterozygosity and hence only a small proportion of cases to be homozygous at the same location. This might be the case if you study many single cases from consanguineous families which might not share the same disaese locus.
The analysis might take several minutes to complete.
The analysis is described in more detail here.
Use autozygosity required only when analysing patients from the same family. Patients from different families will share the same homozygous region when there is no locus heterogeneity but not the same genotypes!
This interface allows you to select one of your own analyses (coloured blue) to inspect homozygosity scores and genotypes. In addition, analyses of other researchers made public (green) or accessible for you (black) will be listed. Analyses are ordered by project name and analysis name. Your own analyses appear on top of the list.
Simply click on the correct analysis to inspect genome-wide homozygosity.
If you are interested in the homozygosity around some candidate genes, use the GeneDistiller link on the right of each item. Please note that GeneDistiller will be opened without any further information and will complain unless you enter your candidate genes or a genomic region of interest.
Here, the homozygosity scores are plotted against the physical position.
The interface can either display the whole genome, a single chromosome or a
selected region on a single chromosome. Interesting regions are indicated by red
bars.
This view becomes more interesting when 'sporadic cases' with a suspected
inbreeding background but without close consanguinity and hence
very short homozygous stretches are studied.
Below the bar chart, you have the possibility to define a specfic region for which the genotypes will be
displayed.
Furthermore, all scores above 60% of the maximum are listed (sorted by their score) and
direct links to these regions are provided. On top of this table, 'broad' regions are displayed -
in these, smaller decreases of the score within a homozygous region are neglected. At the bottom
of the table, 'narrow' regions with sharp limits follow. If you expect some degree of heterozygosity,
you should consider the 'broad' regions.
Below the scores, two links to GeneDistiller are shown, either for the broad or the narrow regions.
Below the regions, links to GeneDistiller and export options are displayed: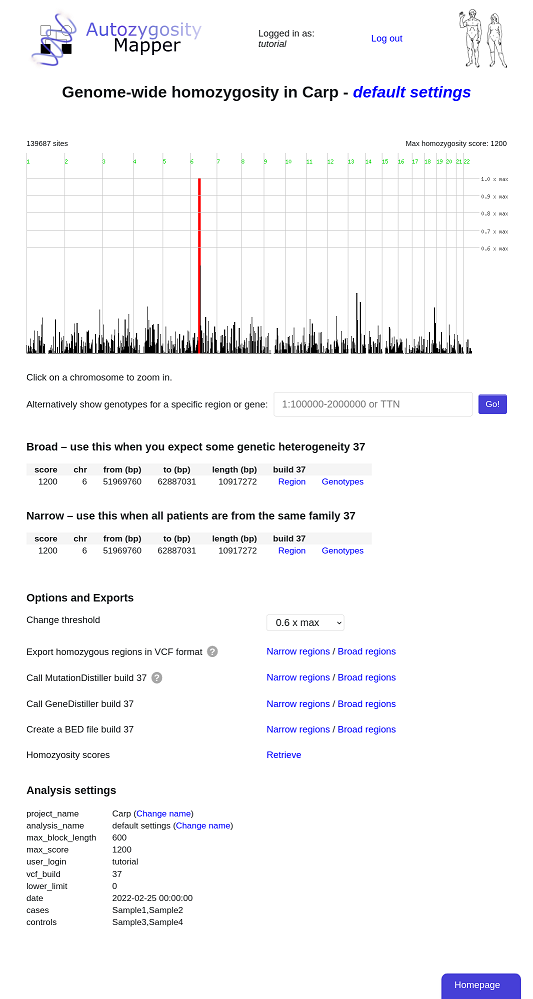
In the whole genome view, clicking on a chromosome will zoom on this chromosome.
After clicking on a chromosome, the view zooms into this chromosome. Now only the interesting regions located within the selected region (here: the chromosome) will be displayes as links.
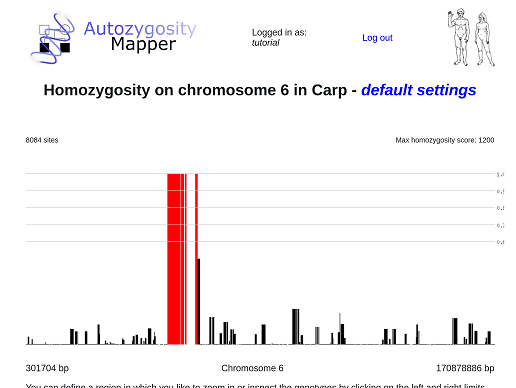
After clicking on the edges of the region the user want to study in details, a pop-up will appear and offer links to further zoom in or to see the underlying genotypes.
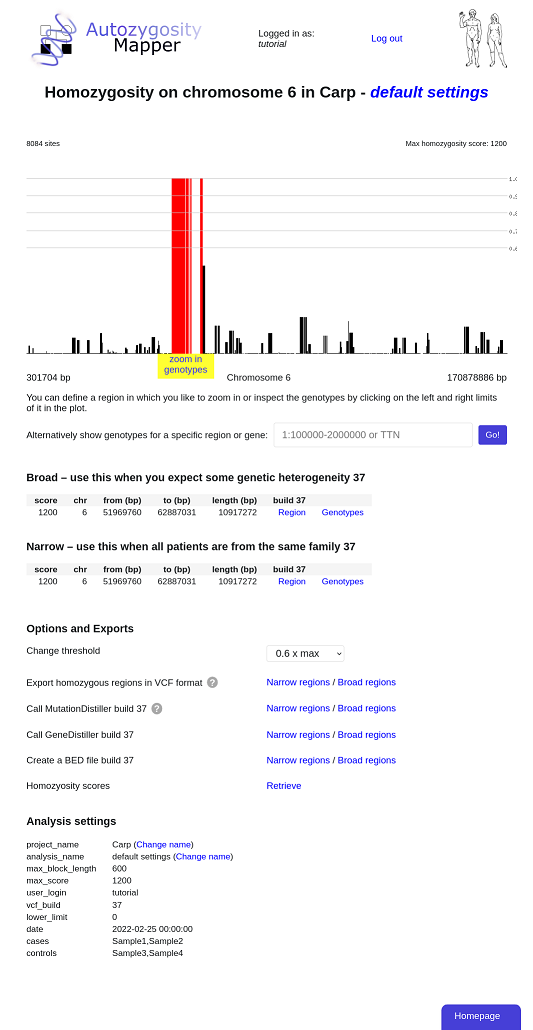
The genotype view depicts the single genotypes of all samples marked as cases or controls. The genotypes are indicated as coloured boxes; where blue codes for heterozygous, grey for unknown and red for homozygous genotypes. Different shades of red are used, longer homozygous stretches will be drawn in a 'deeper' red than single homozygous markers. Genotypes homozygous for another allele are marked with a black diagonal bar.
The homozygous region found by AutozygosityMapper is shown as a rectangle, it's limits can be shifted to other markers by clicking on these.
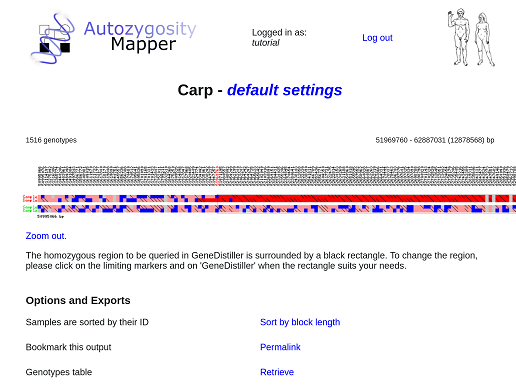
After the region has been defined, you can go ahead by clicking on the GeneDistiller button to find candidate genes (see below) within the region or export only the genotypes within the homozygous or autozygous regions as VCF. You can also directly analyse them in MutationDistiller.
Detailed inspection is possible with the Genotypes table link.
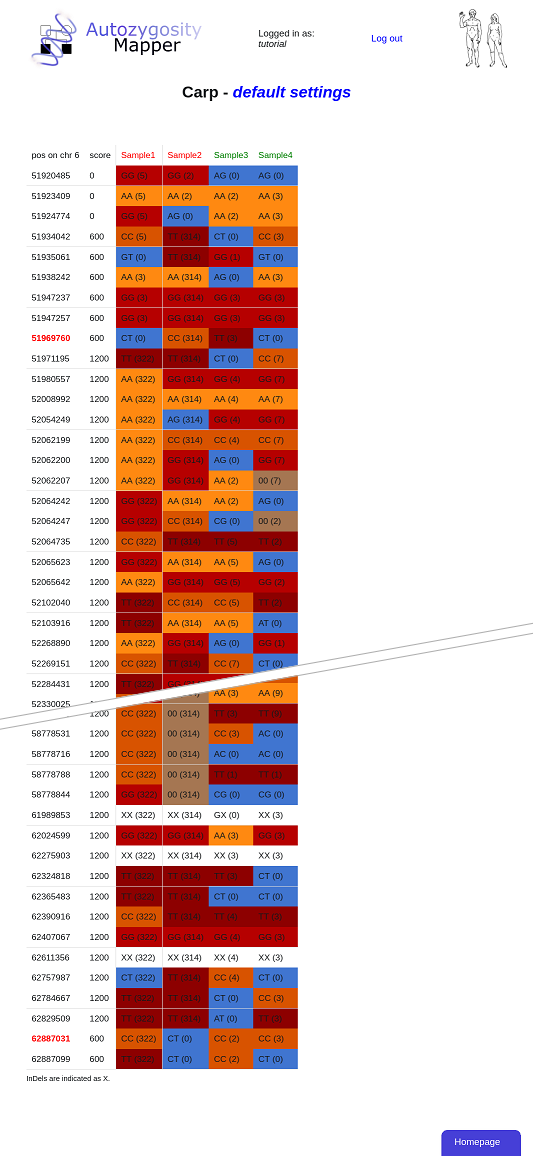
This table shows you the genotypes within the homozygous region. InDels are indicated as X here, but the real alleles are included in the downloadable VCF. The number behind the genotype indicates the length of the homozygous stretch in the individual at this position.
We use different shades of red for the different homozygous alleles - this shows that the region in which Sample1 and Sample2 have the same homozygous genotypes is actually much smaller than their homozygous region. A search for autozygosity instead of homozygosity would have identified this smaller region.
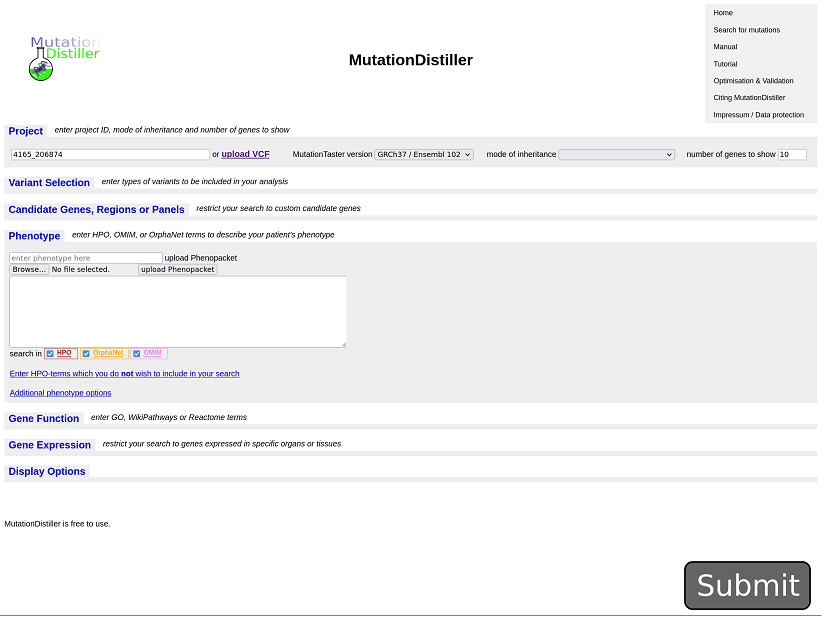
AutozygosityMapper integrates with GeneDistiller. When clicking
the GeneDistiller button, GeneDistiller's query interface will open with the
correct region filled in. Clicking on submit in this interface will list all
genes within the region; for a description how GeneDistiller can be used to focus on genes of interest take a
look at it's manual:
http://www.genedistiller.org/manual.html
When GeneDistiller is queried from AutozygosityMapper, each gene will carry another link, 'AutozygosityMapper', which will open the genotypes view and indicate the position of the selected genes within the genotypes. Bookmarking GeneDistiller's settings (on the bottom of it's results list) will maintain the hyperlinks to the respective homozygosity mapping.
Below the table with the regions you find the possibility to export these regions as a vcf file, which will be generated and downloaded after a click on the respective button. You can choose between the narrow and broad regions.
Below the export option, you find a link to MutationDistiller, a tool for the further analysis of the found regions and identification of pathogenic DNA variants as well as known disease mutations. There, you can upload the exported VCF file with the respective regions and add additional information (if present) for the analysis. A tutorial can be found here.
This analysis results in an overview of genes with potential disease mutations (see pictures below). Further below on the analysis results page, you can find additional information on the gene-linked diseases.


You can use this interface to fine-tune access to your data. It will display a table with all of your projects as rows and all user logins as columns. For each user and project, you can select whether he or she shall be allowed to query your data. You can further grant the right to perform new analyses on your data to collaborators (the project and the analyses will remain under your control though). At the end of each row, you have the option to grant (or revoke) query permissions to the public.
Click on SetPermissions when you're done.
Use this interface to delete single analyses within a project or complete projects.
All of your projects and analyses will be listed. Checking a project/analysis will mark it for deletion; if you delete a project, all analyses performed with that dataset will be deleted as well.
Please note that deleted data will be lost forever.
AutozygosityMapper is based on a PostgreSQL database, all interfaces were written in Perl. The database schema and further details can be found in the technical documentation.
The application was developed using various flavours of Mozilla Firefox. It was further tested with Google Chrome and Microsoft Edge.
We include some benchmarks to demonstrate the speed of AutozygosityMapper. Two examples were uploaded and analysed under normal usage of the webserver and the RDBMS. Please keep in mind that a heavy load on the server and especially a slow network connection may decrease the application's speed significantly.
| Example | Genotypes | SNPs/variants | Samples (cases/controls) | Time (min:s) |
|
| SNP tutorial genotypes | Affymetrix 50K Hind 240 (download) |
56,936 | 8 (5/3) |
upload & homozygosity detection | 1:08 |
| analysis | 0:06 | ||||
| display of genome-wide homozygosity | 0:01 | ||||
| VCF tutorial genotypes | WES VCF (download) |
139,733 | 4 (2/2) | upload & homozygosity detection |
0:21
|
| analysis (homozygosity) | 0:06 | ||||
| analysis (autozygosity) | 0:06 | ||||
| display of genome-wide homozygosity | 0:01 | ||||
| VCF WES genotypes | very large WGS VCF (download) |
17,740,640 | 4 (2/2) | upload & homozygosity detection |
58:42
|
| analysis (homozygosity) | 5:56 | ||||
| analysis (autozygosity) | 5:45 | ||||
| display of genome-wide homozygosity | 0:32 |
In addition to the web-based version, a stand-alone application exists. This program does not make use of a database and interfaces were created with Perl/Tk. While it offers speed gains for small chips (up to 50,000 SNPs), the performance decreases with the number of markers being studied. This version is not supported anymore but we will gladly share the source code. Please send us an e-mail if you are interested.
| Dominik Seelow | Berliner Institut für Gesundheitsforschung | Charité, Berlin, Germany |
| Robin Steinhaus | Berliner Institut für Gesundheitsforschung | Charité, Berlin, Germany |
| Melanie Vogel | Freie Universität Berlin, Berlin, Germany |
| Björn Fischer-Zirnsak | Charité, Berlin, Germany |
| Felix Boschann | Charité, Berlin, Germany |
Email: dominik.seelow(at)charite.de
If you discovered bugs or have suggestion, please write us an email to our ticketing system.
Why is this software still using genome build 37?
Because it is still the most-widely used genome version in medical genetics. Besides, the SNP array
annotation files refer to 37.
We will certainly offer build 38 in the future.
Do I have access to my HomozygosityMapper projects on
AutozygosityMapper?
No.
Why do my results differ between HomozygosityMapper and
AutozygosityMapper?
HM uses the number of consecutive homozygous genotypes to calculate the length of homozygous segments,
AM uses their physical length. The first approach was perfect for SNP arrays with evenly-distributed
markers
but it performs worse with WES genotypes where only protein-coding genes are covered. A large number of
homozygous
variants within one gene gives a very high score in HM, even if the homozygous stretch is relatively
short.
Will my HomozygosityMapper projects remain?
Yes, as long as you created them while logged in (please delete your projects as soon as possible!).
Do I have to create a new account?
We took over all HomozygosityMapper accounts by Dec 15th 2020. Accounts that were created after that
date are either valid for HomozygosityMapper OR AutozygosityMapper.
Why am I not able to find the allele frequencies anymore?
For now, they are not required for the search for homozygous regions. Furthermore, they would require
more storage space and especially a higher run time.
Can I archive projects in HomozygosityMapper and restore them in
AutozygosityMapper?
No, because the structure of the data base has changed considerably.
Why can't I further zoom in on a single chromosome? Why can't I change the limits of
a homozygous region?
Sounds as if JavaScript was not enabled. With Mozilla Firefox, select Tools -> Options -> Content
and check the 'Enable JavaScript' button.
I forgot my login and/or my password. What can I do?
Please recover your credentials via the "I forgot my credentials!" link on the login page. Your data
will be sent to you via the e-mail address you enter here, in case this is the address you registered
your account with.
I am using a chip that is not listed, how can I upload my genotypes?
So far, AutozygosityMapper only supports the most common chips manufactured by Affymetrix and
Illumina. If you want to import genotypes derived by other chips (or means), you can either bring your
genotype file in the format as used by Affymetrix or Illumina and select a chip containing all your
markers or write us an e-mail. The integration of completely new chips and file formats might take some
time, though.
I made a terrible mistake and deleted my project. Is there any way to restore
it?
Well, yes, at least there might be one. If you are lucky, the data is included in one of our backups.
Please write an e-mail to us and specify the exact name of the project. We will then see what we can do.
Is the data within AutozygosityMapper stored forever?
Hm. Certainly not forever but at least we do not plan to include an automatic time-out after which your
data will be deleted. We might, for disk space reasons, decide to archive 'old projects'. However, we
will inform the respective project's owners in advance.
I have created an account but I cannot log in. Why?
Please ensure that you allowed this site to set cookies in your browser. This is the default setting in
all current browsers - if you deactivated cookies you could change the setting to 'ask' instead of
'block' or add an exception for this website.
How is this homozgyosity score calculated?
AM calculates the length of the homozygous block (in SNPs) at each position/marker for each sample.
The values of the 'cases' are then added to get the 'homozygosity score' for a position.
It is, however, actually a bit more complex: The maximum length for each block (that will be
added to the final score) is per default set to some limits (depending on the user-defined
'limit block length' or default value) to reduce the effect of very long blocks in one or few samples.
That means, that a very long block (say 100 mbp) is 'reduced' to the 'limit block length'
setting (say 6 mbp).
For case-control studies or 'distant consanguinity', only blocks longer than a user-defined
value 'only count blocks longer than' are regarded to reduce the influence of non-informative
segments.
And, of course, in the case of 'genetic homogeneity', regions are excluded
when the same homozygous haplotype is found in any controls.
The analysis is described in more detail in the technical documentation.
Why are your sample VCF so much smaller than my own VCFs?
We stripped them of all data not required to use AutozygosityMapper.
Species pictures
Cow: LadyofHats
Dog: Mathieu19
Humans: L. Salzman Sagan, C.
Sagan, F. Drake / NASA
Horse: Wilfredor
Mouse: The Tango! Desktop Project
Rat: Martin Krzywinsk
Sheep: Michał Pecyna